Docking Proteins to Deny Disease: Computational Considerations for Simulating Protein-Ligand Interaction | Exxact Blog

Benchmarking the performance of irregular computations in AutoDock-GPU molecular docking - ScienceDirect
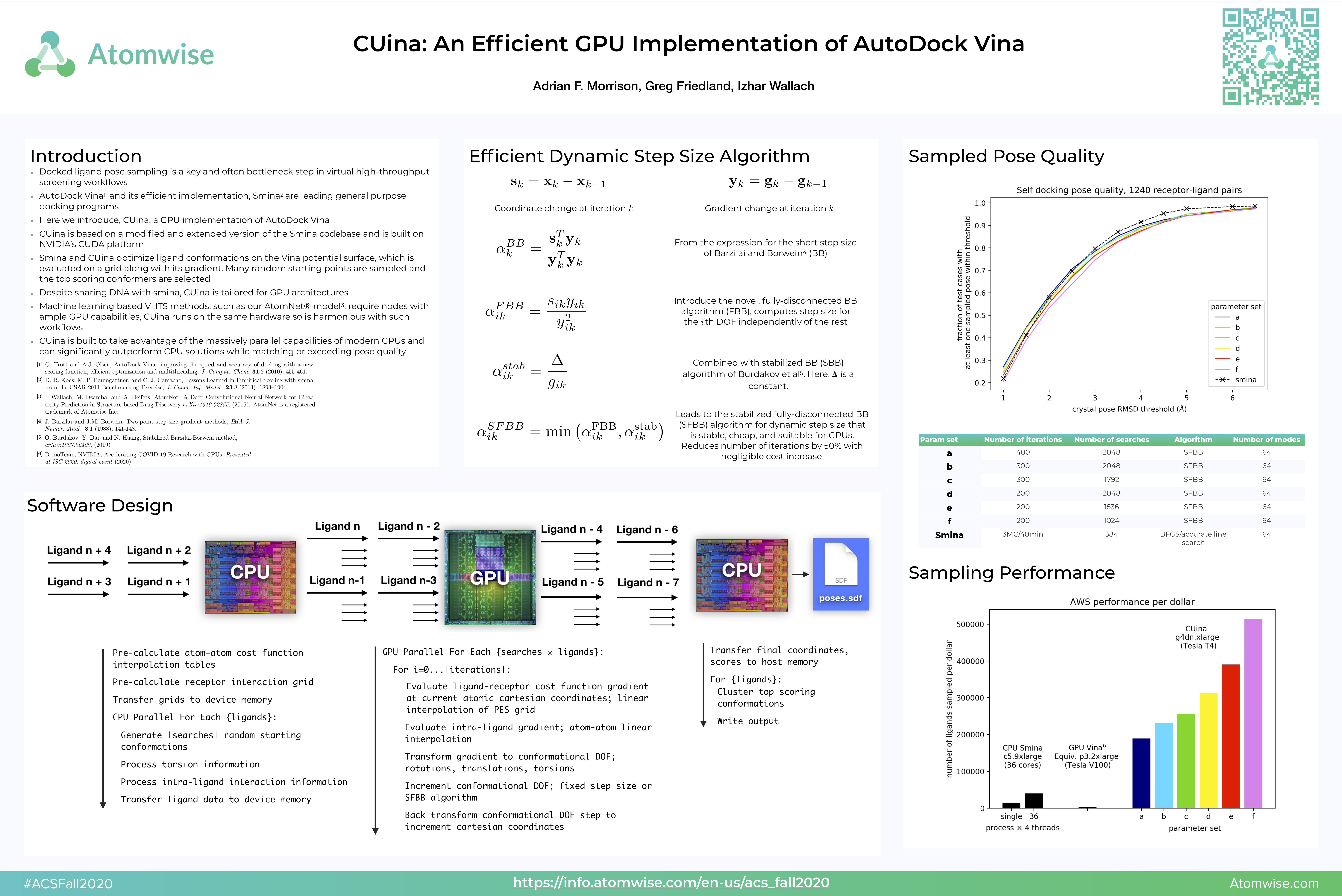
CUina: An Efficient Implementation of Autodock Vina Specially Crafted for CUDA GPUs | NVIDIA On-Demand

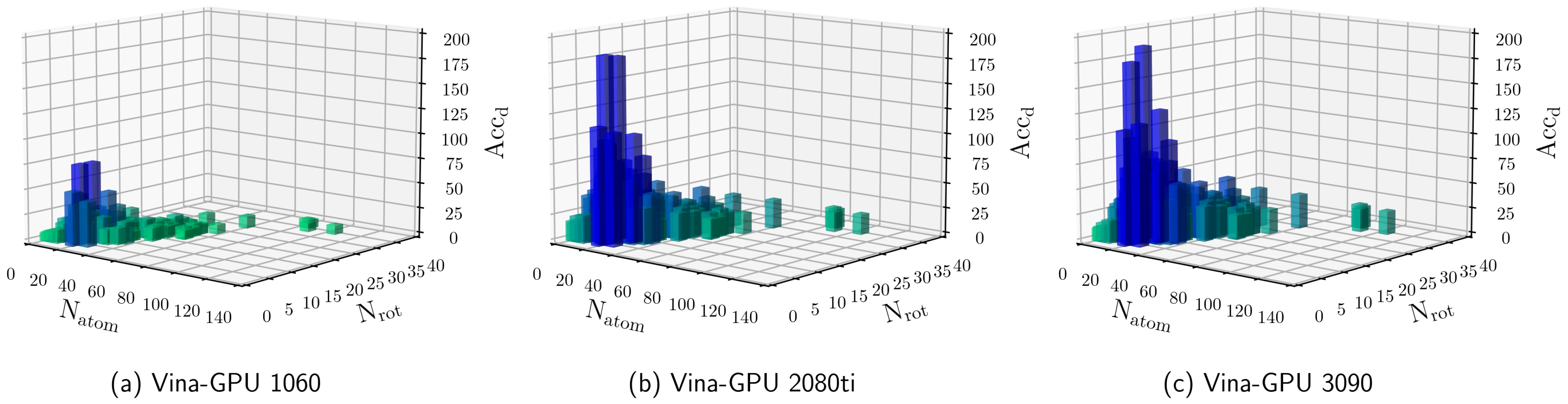
Docking scores of VINA-GPU on six PDB structures with various levels of... | Download Scientific Diagram

Benchmarking the performance of irregular computations in AutoDock-GPU molecular docking - ScienceDirect
GitHub - purnawanpp/Docking-4ieh: Docking Tutorial Using Autodock Vina version 1.2.3 (2021) and AutoDock-GPU Version 1.5.3